I have a problem with combining ggplots using cowplot and ggpubr which is driving me crazy.
The problem is with the legend. When it's displayed with its original graph on its own, the spacing between the elements of the legend (guide title, key, key-label) are absolutely fine.
However, if I extract the legend from the original plot, and then display it in a combined plot (using either plot_grid from cowplot or ggarrange from ggpubr) then the spacing goes haywire. The longer the text, the more the spacing expands.
What is wrong here, and how do I fix it so that the legend in the combined plot looks exactly like the one in the original individual plot?
Example
This example uses ggarrange from ggpubr; my results with get_legend and plot_grid using cowplot are similar. Treatment names are entirely made up.
library(survival)
library(broom)
library(dplyr)
library(foreach)
library(ggpubr)
fit <- survfit(Surv(time,status == 2) ~ trt + sex, data=pbc)
time.xticks <- seq(0, 4000, 1000)
delta <- 0.00001
# Survival plot
kmdata <- tidy(fit) %>%
mutate(trt=factor(gsub('trt=(.*),.*','\\1',strata)),
sex=factor(gsub('.*sex=(.*)','\\1',strata), levels=levels(pbc$sex)))
p1 <- ggplot(data=filter(kmdata, time<=max(time.xticks)), aes(x=time, y=estimate, colour=sex, linetype=trt)) + geom_step() +
scale_x_continuous(breaks = time.xticks,
limits = c(min(time.xticks), max(time.xticks))) +
scale_colour_discrete(name="Sex", labels=c("Male","Female")) +
scale_linetype_discrete(name="Treatment group", labels=c("Zyxatrxilbroh 35 mg", "Placebo 35 mg")) +
theme(legend.position = "bottom", legend.box = "horizontal",
legend.background = element_rect(fill="grey90", colour="black", size=0),
legend.key.height=unit(0.2, "cm"),
text=element_text(size=18))
tardata <- foreach(s=unique(kmdata$strata), .combine="rbind") %do% {
filter(kmdata, strata==s)[findInterval(pmax(0, time.xticks-delta), filter(kmdata, strata==s)$time)+1,] %>%
bind_cols(tibble(time.xticks))
} %>%
mutate(ypos = -((as.integer(sex)-1)*(length(unique(pbc$trt))+2) + as.integer(trt) + 1))
tarheads <- tibble(xpos=0,
ypos=-(((1:length(unique(pbc$sex))) - 1)*(length(unique(pbc$trt)) + 2) + 1),
lab=levels(pbc$sex))
risk.yticks <- sort(unique(tardata$ypos))
risk.ylabels <- rep(rev(paste("trt =",levels(kmdata$trt))), length(unique(kmdata$sex)))
# Number-at-risk table
p2 <- ggplot(data=tardata, aes(x=time.xticks, y=ypos, label=n.risk, colour=sex)) + geom_text() +
geom_text(data=tarheads, aes(x=xpos, y=ypos, label=lab), colour="black", hjust="left") +
scale_x_continuous(breaks = time.xticks,
limits = c(min(time.xticks), max(time.xticks))) +
scale_y_continuous(breaks = risk.yticks,
labels = risk.ylabels) +
theme(text=element_text(size=18))
# put the two together
p.comb <- ggarrange(p1, p2, heights = c(2, 0.8), ncol=1,
align="v", common.legend = TRUE, legend="bottom")
# alternate version with guide headings at the top left instead of at the side
p1.a <- p1 + guides(colour = guide_legend(order=1,
title.position = "top",
title.hjust = 0),
linetype = guide_legend(order=1,
title.position = "top",
title.hjust = 0))
p.comb.a <- ggarrange(p1.a, p2, heights = c(2, 0.8), ncol=1,
align="v", common.legend = TRUE, legend="bottom")
# send to png
png("test-p1.png", width=8, height=4.5, units="in", res=200, type="cairo")
plot(p1)
dev.off()
png("test-pcomb.png", width=8, height=4.5, units="in", res=200, type="cairo")
plot(p.comb)
dev.off()
Results
Individual plot with correct legend spacing:
[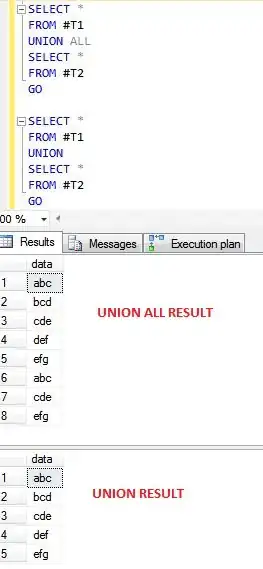
Combined plot with legend spacing expanded so much that the legend no longer fits in the image: