I have a some data and want to fit a given psychometric function p.
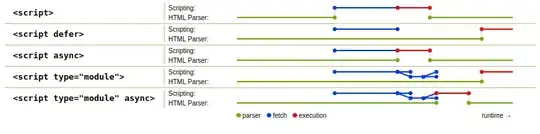 I'm intereseted in the fit parameters and the errors as well. With the 'classical' method using the curve_fit function from the scipy package it's easy to get the parameters of p and the errors. However I want to do the same using a maximum likelihood estimation (MLE). From the output and the figure you can see that both methods offer slight different parameters. Implementing the MLE is not the problem but I don't know how to get the errors using this method. Is there an easy way to get them? My likelihood function L is:
I'm intereseted in the fit parameters and the errors as well. With the 'classical' method using the curve_fit function from the scipy package it's easy to get the parameters of p and the errors. However I want to do the same using a maximum likelihood estimation (MLE). From the output and the figure you can see that both methods offer slight different parameters. Implementing the MLE is not the problem but I don't know how to get the errors using this method. Is there an easy way to get them? My likelihood function L is:
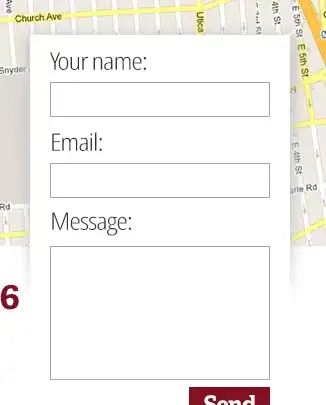 I was not able to adapt the code described here http://rlhick.people.wm.edu/posts/estimating-custom-mle.html but this is probably a solution. How can I implement this? Or this there any other way?
I was not able to adapt the code described here http://rlhick.people.wm.edu/posts/estimating-custom-mle.html but this is probably a solution. How can I implement this? Or this there any other way?
A similar function is fitted here using scipy stats models: https://stats.stackexchange.com/questions/66199/maximum-likelihood-curve-model-fitting-in-python. However the errors of the parameters are not calculated neither.
The negative log-likelihood function is correct, since it offers the right parameters, but I was wondering if this function depends on y-data? The negative log likelihood function l is obviously l = -ln(L). Here is my code:
#!/usr/bin/env python
# -*- coding: utf-8 -*-
## libary
import numpy as np
import matplotlib.pyplot as plt
from scipy.optimize import curve_fit
from scipy.optimize import minimize
def p(x,x50,s50):
"""return y value of psychometric function p"""
return 1./(1+np.exp(4.*s50*(x50-x)))
def initialparams(x,y):
"""return initial fit parameters for function p with given dataset"""
midpoint = np.mean(x)
slope = (np.max(y)-np.min(y))/(np.max(x)-np.min(x))
return [midpoint, slope]
def cfit_error(pcov):
"""return errors of fir from covariance matrix"""
return np.sqrt(np.diag(pcov))
def neg_loglike(params):
"""analytical negative log likelihood function. This function is dependend on the dataset (x and y) and the two parameters x50 and s50."""
x50 = params[0]
s50 = params[1]
i = len(xdata)
prod = 1.
for i in range(i):
#print prod
prod *= p(xdata[i],x50,s50)**(ydata[i]*5) * (1-p(xdata[i],x50,s50))**((1.-ydata[i])*5)
return -np.log(prod)
xdata = [0.,-7.5,-9.,-13.500001,-12.436171,-16.208617,-13.533123,-12.998025,-13.377527,-12.570075,-13.320075,-13.070075,-11.820075,-12.070075,-12.820075,-13.070075,-12.320075,-12.570075,-11.320075,-12.070075]
ydata = [1.,0.6,0.8,0.4,1.,0.,0.4,0.6,0.2,0.8,0.4,0.,0.6,0.8,0.6,0.2,0.6,0.,0.8,0.6]
intparams = initialparams(xdata, ydata)## guess some initial parameters
## normal curve fit using least squares algorithm
popt, pcov = curve_fit(p, xdata, ydata, p0=intparams)
print('scipy.optimize.curve_fit:')
print('x50 = {:f} +- {:f}'.format(popt[0], cfit_error(pcov)[0]))
print('s50 = {:f} +- {:f}\n'.format(popt[1], cfit_error(pcov)[1]))
## fitting using maximum likelihood estimation
results = minimize(neg_loglike, initialparams(xdata,ydata), method='Nelder-Mead')
print('MLE with self defined likelihood-function:')
print('x50 = {:f}'.format(results.x[0]))
print('s50 = {:f}'.format(results.x[1]))
#print results
## ploting the data and results
xfit = np.arange(-20,1,0.1)
fig = plt.figure()
ax = fig.add_subplot(1,1,1)
ax.plot(xdata, ydata, 'xb', label='measured data')
ax.plot(xfit, p(xfit, *popt), '-r', label='curve fit')
ax.plot(xfit, p(xfit, *results.x), '-g', label='MLE')
plt.legend()
plt.show()
The output is:
scipy.optimize.curve_fit:
x50 = -12.681586 +- 0.252561
s50 = 0.264371 +- 0.117911
MLE with self defined likelihood-function:
x50 = -12.406544
s50 = 0.107389
Both fits and measured data can be seen here:
 My Python version is 2.7 on Debian Stretch. Thank you for your help.
My Python version is 2.7 on Debian Stretch. Thank you for your help.