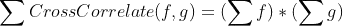I am trying to compare several datasets and basically test, if they show the same feature, although this feature might be shifted, reversed or attenuated. A very simple example below:
A = np.array([0., 0, 0, 1., 2., 3., 4., 3, 2, 1, 0, 0, 0])
B = np.array([0., 0, 0, 0, 0, 1, 2., 3., 4, 3, 2, 1, 0])
C = np.array([0., 0, 0, 1, 1.5, 2, 1.5, 1, 0, 0, 0, 0, 0])
D = np.array([0., 0, 0, 0, 0, -2, -4, -2, 0, 0, 0, 0, 0])
x = np.arange(0,len(A),1)
I thought the best way to do it would be to normalize these signals and get absolute values (their attenuation is not important for me at this stage, I am interested in the position... but I might be wrong, so I will welcome thoughts about this concept too) and calculate the area where they overlap. I am following up on this answer - the solution looked very elegant and simple, but I may be implementing it wrongly.
def normalize(sig):
#ns = sig/max(np.abs(sig))
ns = sig/sum(sig)
return ns
a = normalize(A)
b = normalize(B)
c = normalize(C)
d = normalize(D)
But then, when I try to implement the solution from the answer, I run into problems.
OLD
for c1,w1 in enumerate([a,b,c,d]):
for c2,w2 in enumerate([a,b,c,d]):
w1 = np.abs(w1)
w2 = np.abs(w2)
M[c1,c2] = integrate.trapz(min(np.abs(w2).any(),np.abs(w1).any()))
print M
Produces TypeError: 'numpy.bool_' object is not iterable or IndexError: list assignment index out of range. But I only included the .any() because without them, I was getting the ValueError: The truth value of an array with more than one element is ambiguous. Use a.any() or a.all().
EDIT - NEW (thanks @Kody King)
The new code is now:
M = np.zeros([4,4])
SH = np.zeros([4,4])
for c1,w1 in enumerate([a,b,c,d]):
for c2,w2 in enumerate([a,b,c,d]):
crossCorrelation = np.correlate(w1,w2, 'full')
bestShift = np.argmax(crossCorrelation)
# This reverses the effect of the padding.
actualShift = bestShift - len(w2) + 1
similarity = crossCorrelation[bestShift]
M[c1,c2] = similarity
SH[c1,c2] = actualShift
M = M/M.max()
print M, '\n', SH
And the output:
[[ 1. 1. 0.95454545 0.63636364]
[ 1. 1. 0.95454545 0.63636364]
[ 0.95454545 0.95454545 0.95454545 0.63636364]
[ 0.63636364 0.63636364 0.63636364 0.54545455]]
[[ 0. -2. 1. 0.]
[ 2. 0. 3. 2.]
[-1. -3. 0. -1.]
[ 0. -2. 1. 0.]]
The matrix of shifts looks ok now, but the actual correlation matrix does not. I am really puzzled by the fact that the lowest correlation value is for correlating d with itself. What I would like to achieve now is that:
EDIT - UPDATE
Following on the advice, I used the recommended normalization formula (dividing the signal by its sum), but the problem wasn't solved, just reversed. Now the correlation of d with d is 1, but all the other signals don't correlate with themselves.
New output:
[[ 0.45833333 0.45833333 0.5 0.58333333]
[ 0.45833333 0.45833333 0.5 0.58333333]
[ 0.5 0.5 0.57142857 0.66666667]
[ 0.58333333 0.58333333 0.66666667 1. ]]
[[ 0. -2. 1. 0.]
[ 2. 0. 3. 2.]
[-1. -3. 0. -1.]
[ 0. -2. 1. 0.]]
- The correlation value should be highest for correlating a signal with itself (i.e. to have the highest values on the main diagonal).
- To get the correlation values in the range between 0 and 1, so as a result, I would have 1s on the main diagonal and other numbers (0.x) elsewhere.
I was hoping the M = M/M.max() would do the job, but only if condition no. 1 is fulfilled, which it currently isn't.